Anvil helps develop new SARS-CoV-2 therapy
A research team from Delaware State University used Purdue’s Anvil supercomputer to computationally design a new pan-coronavirus therapy that is resistant to mutational changes.
Dr. Mohammad Shahidul Islam, a 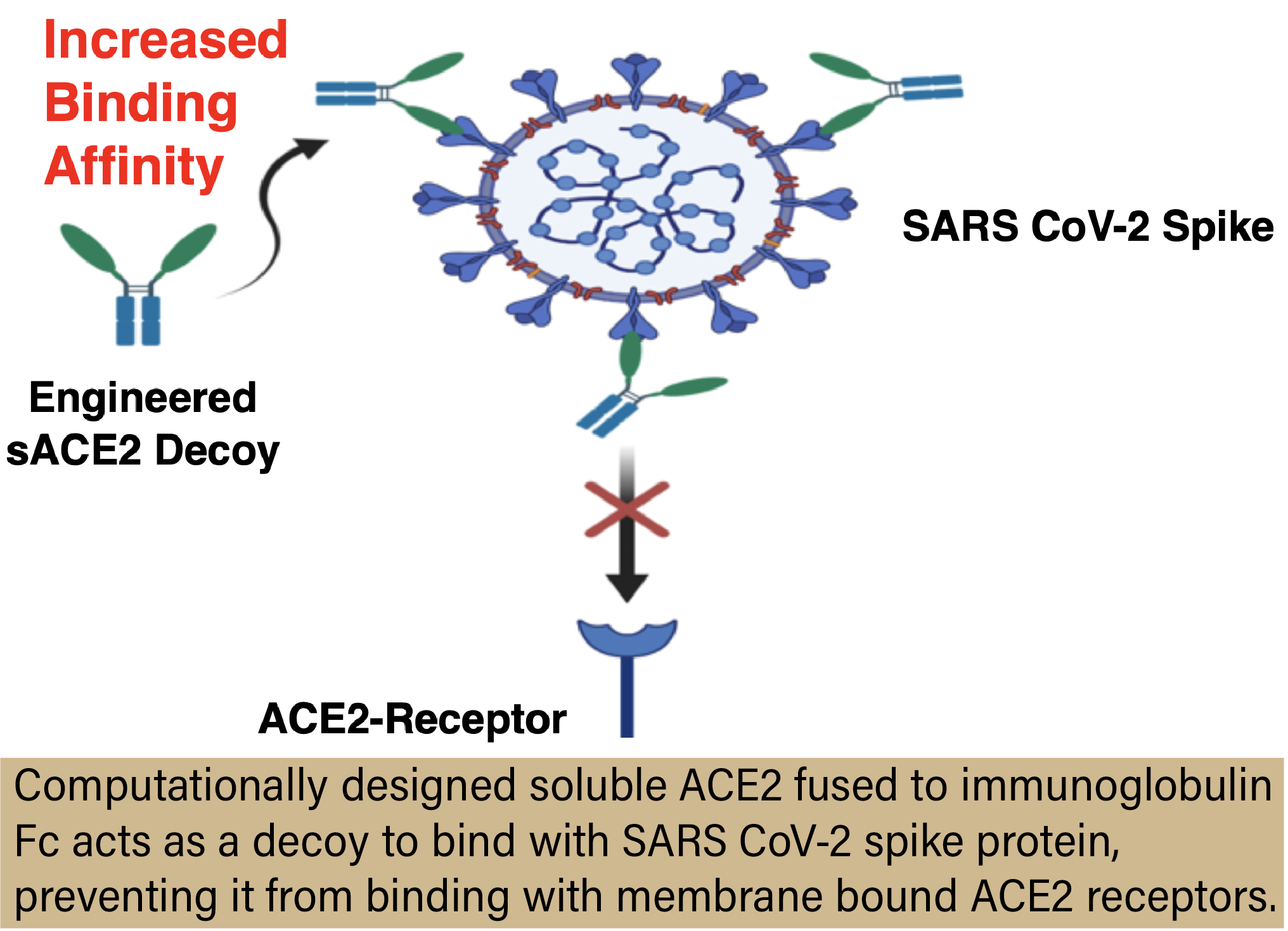 former assistant professor in the Department of Chemistry at the University of Illinois Chicago (UIC) and current associate professor at Delaware State University, and Brandon Havranek, a former research assistant at UIC and current medical student at Thomas Jefferson University, conducted research into creating a better, more resilient, and more effective therapy for severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), the causative agent of COVID-19 disease. The team succeeded in computationally engineering a soluble ACE2 decoy receptor that proved to be effective both in vitro and in vivo. While the ACE2 decoy is not yet available for use in humans, this research helps to showcase that computational methods are accurate enough to design therapeutics for viruses.
former assistant professor in the Department of Chemistry at the University of Illinois Chicago (UIC) and current associate professor at Delaware State University, and Brandon Havranek, a former research assistant at UIC and current medical student at Thomas Jefferson University, conducted research into creating a better, more resilient, and more effective therapy for severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), the causative agent of COVID-19 disease. The team succeeded in computationally engineering a soluble ACE2 decoy receptor that proved to be effective both in vitro and in vivo. While the ACE2 decoy is not yet available for use in humans, this research helps to showcase that computational methods are accurate enough to design therapeutics for viruses.
SARS-CoV-2 infects our bodies by binding to the ACE2 receptors on cell membranes. These ACE2 receptors are ubiquitous throughout the body but are found in abundance in certain areas, like the lungs. Once the virus binds to the receptor, it can enter the cell and begin to replicate. This is how the virus is able to survive after entering the system.
Since the COVID-19 pandemic began, many therapeutics have been developed with the goal of blocking SARS-CoV-2 from attaching to the ACE2 receptor, specifically, protein minibinders, peptides, monoclonal antibodies, and nanobodies. All of these therapies, however, face an issue that can render them ineffective—genetic mutation. Most of these therapeutics are developed against the original SARS-CoV-2 virus or a specific variant, such as delta or omicron, and can be very effective in neutralizing the particular strain for which it was made. The problem is in how quickly SARS-CoV-2 mutates. Once a genetic mutation occurs, the therapeutic of choice no longer recognizes the virus and does not block the interaction. And as we have seen, SARS-CoV-2 mutates quite rapidly, making it very difficult to contain or eradicate.
The solution, then, is to create a pan-coronavirus therapy that does not rely on targeting specific genetic strains of the virus, which is precisely what the team from UIC set out to do.
The group used the Anvil supercomputer to computationally develop an affinity-enhanced soluble ACE2 decoy receptor, known as FLIF. The goal of a soluble ACE2 decoy receptor is exactly what it sounds like—to act as a decoy. Administered to the bloodstream via nasal spray or through an IV, the ACE2 decoy floats around as an alternative landing site for the virus. In theory, when the virus enters our system, it will attach to the ACE2 decoy receptors instead of the natural ACE2 receptors on our cell membranes, keeping us from becoming infected and effectively neutralizing the virus. Dr. Islam and Havranek are not the first researchers to pursue this type of therapy. But the inherent problem thus far has been that the binding affinity for soluble ACE2 decoy receptors is low; i.e. the virus doesn’t like them and still attaches to our cells. Where FLIF excels is in the fact that it was computationally developed to be affinity-enhanced, or more attractive, for SARS-CoV-2. FLIF outcompetes our native ACE2 receptors, making it highly effective as a therapeutic.
“The affinity of soluble ACE2 essentially wasn’t strong enough to intercept a lot of the spike proteins entering the body, so a lot of them could still bind to the cells and lead to infection,” says Havranek. “What we needed to do was to engineer the soluble ACE2 decoy to have higher affinity than the membrane-bound protein, so that it could outcompete it essentially. And we did that computationally using the Anvil supercomputer.”
Using software called Rosetta, Havranek and Dr. Islam designed FLIF, a 4-mutation soluble ACE2 decoy that proved to have a roughly 80-fold tighter binding affinity (for the delta variant) than non-affinity enhanced soluble ACE2. The process of designing FLIF was extensive, requiring the group to test multiple variations of mutations to see if any improved the affinity to SARS-CoV-2 via binding free-energy calculations.
“So with Anvil,” says Havranek, “we could basically test out a number of different mutations, which would be really hard to do on a normal computer. It’d probably take months. But on Anvil, we could do that testing relatively quickly. And so we came up with a number of different mutations—four to be exact. And then, what we did was, we did computational binding free energy calculations, which would also take a very long time on just a normal computer. And we tested our design against the soluble version that had no mutations against a bunch of different variants of SARS-CoV-2.”
The team tested FLIF for the SARS-CoV-2 delta variant, multiple omicron variants, SARS-CoV-1, and a number of sarbecoviruses that have yet to cross over from animals to humans but are suspected to be potential threats. FLIF proved to be highly effective against all types, and not just computationally.
Dr. Islam and Havranek wanted to validate their computational results, so they collaborated with experimental researchers across multiple institutions. The team tested the efficacy of FLIF using a multitude of binding experiments, including flow cytometry and biolayer interferometry. FLIF showed the ability to neutralize all variants tested. The team also tested FLIF against a non-affinity-enhanced ACE2 decoy in Syrian hamsters and found that FLIF was effective in both neutralizing the virus and lowering cytokine production, whereas the non-enhanced ACE2 was not.
“According to our investigation,” says Dr. Islam, “we think that the FLIF decoy ACE2 that we developed is currently [as of June 2023] the best computationally designed decoy ACE2 out there. So with Anvil and access to other supercomputers, we were able to get that done.”
The results from Dr. Islam, Havranek, and the experimental team’s research are nothing short of extraordinary. The only way in which FLIF could be rendered ineffective would be a mutation in the RBD (receptor-binding domain) of the spike protein in SARS-CoV-2 that limits binding to the ACE2 decoy. But, according to Havranek, any such mutation would also reduce binding to native ACE2 receptors, making the virus unfit to propagate, so it is not a major concern. Overall, it seems that affinity-enhanced ACE2 decoys will be necessary to fight against evolving SARS-CoV-2 variants, and continued research into computationally designing these therapeutics is of the utmost importance.
The collaborative experimental research team was composed of the following individuals:
-Graeme Walker Lindsey, Department of Biochemistry, University of Illinois
-Yusuke Higuchi, Department of Cardiovascular Medicine, Graduate School of Medical Science, Kyoto Prefectural University of Medicine
-Yumi Itoh, Institute for Advanced Co-Creation Studies, Research Institute for Microbial Diseases, Osaka University
-Tatsuya Suzuki, Institute for Advanced Co-Creation Studies, Research Institute for Microbial Diseases, Osaka University
-Toru Okamoto, Institute for Advanced Co-Creation Studies, Research Institute for Microbial Diseases, Osaka University
-Atsushi Hoshino, Department of Cardiovascular Medicine, Graduate School of Medical Science, Kyoto Prefectural University of Medicine
-Erik Procko, Department of Biochemistry, University of Illinois
To learn more about the team’s research, please refer to their Nature Communications Biology publication: A computationally designed ACE2 decoy has broad efficacy against SARS-CoV-2 omicron variants and related viruses in vitro and in vivo
Anvil is Purdue University’s most powerful supercomputer, providing researchers from diverse backgrounds with advanced computing capabilities. Built through a $10 million system acquisition grant from the National Science Foundation (NSF), Anvil supports scientific discovery by providing resources through the NSF’s Advanced Cyberinfrastructure Coordination Ecosystem: Services & Support (ACCESS), a program that serves tens of thousands of researchers across the United States.
For more information regarding HPC and how it can help you, please visit our “Why HPC?” page.
Researchers may request access to Anvil via the ACCESS allocations process. More information about Anvil is available on Purdue’s Anvil website. Anyone with questions should contact anvil@purdue.edu. Anvil is funded under NSF award No. 2005632.
Written by: Jonathan Poole, poole43@purdue.edu
